Gene is usually regarded as the smallest physical unit of heredity encoding a molecular cell product.
G. J. Mendel (1866) was the first to visualize it as a unit of inheritance and called it an element (atoms of heredity), which was termed as a factor by Carl Correns (1900). The term ‘gene’ was coined by W.Johannsen (1909) for a single unit of heredity occupying a specific position (locus) in a chromosome and taking part in expression of a trait. In 1903, W. S. Sutton pointed out that the segregation and recombination of Mendelian factors showed a parallelism in the behaviour of chromosomes. T.H. Morgan (1910–1920) regarded chromosomes as group of linked genes (loci). He found that alternative forms of genes (alleles) at a locus could mutate from one to another.
The importance of genes in enzyme production first emerged through the work on the chemistry and inheritance of eye colour of Drosophila. Through the work on auxotrophic mutants of the mould Neurospora crassa,G.W. Beadle and E.L. Tatum (1948) proposed one gene- one enzyme hypothesis, and considered genes as the heritable determinants of enzymes. Seymour Benzer (1955) recognized three units of genes-cistron (functional unit), recon (recombinational unit) and muton (mutational).
- Deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) are the two types of nucleic acids found in living systems. DNA acts as the genetic material in most of the organisms.
- RNA though it also acts as a genetic material in some viruses, mostly functions as a messenger. RNA has additional roles as well. It functions as adapter, structural, and in some cases as a catalytic molecule.
- In this topic we are going to discuss the structure of DNA, its replication, the process of making RNA from DNA (transcription), the genetic code that determines the sequences of amino acids in proteins, the process of protein synthesis (translation) and elementary basis of their regulation. The determinationof complete nucleotide sequence of human genome during last decade has set in a new era of genomics. In the last section, the essentials of human genome sequencing and its consequences will also be discussed.
Also Check: Human Health
DNA
DNA is a long polymer of deoxyribonucleotides. The length of DNA is usually defined as number of nucleotides (or a pair of nucleotide referred to as base pairs) present in it. This also is the characteristic of an organism. For example, a bacteriophage known as 174 has 5386 nucleotides, Bacteriophage has 48502 base pairs (bp), Escherichia coli has 4.6 × 106 bp, and haploid content of human DNA is
3.3 × 109 bp.
Structure of Polynucleotide Chain
- A nucleotide has three components – a nitrogenous base, a pentose sugar (ribose in case of RNA, and deoxyribose for DNA), and a phosphate group.
- There are two types of nitrogenous bases – Purines (Adenine and Guanine), and Pyrimidines (Cytosine, Uracil and Thymine). Cytosine is common for both DNA and RNA and Thymine is present in DNA. Uracil is present in RNA at the place of Thymine.
- A nitrogenous base is linked to the pentose sugar through a N-glycosidic linkage to form a nucleoside, such as adenosine or deoxyadenosine, guanosine or deoxyguanosine, cytidine or deoxycytidine and uridine or deoxythymidine.
- When a phosphate group is linked to 5'-OH of a nucleoside through phosphoester linkage, a corresponding nucleotide (or deoxynucleotide depending upon the type of sugar present) is formed. Two nucleotides are linked through 3'-5' phosphodiester linkage to form a dinucleotide. More nucleotides can be joined in such a manner to form a polynucleotide chain.
- A polymer thus formed has at one end a free phosphate moiety at
A Polynucleotide chain
5'-end of ribose sugar, which is referred to as 5’-end of polynucleotide chain. Similarly, at the other end of the polymer the ribose has a free 3'-OH group which is referred to as 3' -end of the polynucleotide chain. The backbone in a polynucleotide chain is formed due to sugar and phosphates. The nitrogenous bases linked to sugar moiety project from the backbone (Figure 6.1).
- In RNA, every nucleotide residue has an additional –OH group present at 2' -position in the ribose. Also, in RNA the uracil is found at the place of thymine (5-methyl uracil, another chemical name for thymine).
- DNA as an acidic substance present in nucleus was first identified by Friedrich Meischer in 1869. He named it as ‘Nuclein’. However, due to technical limitation in isolating such a long polymer intact, the elucidation of structure of DNA remained elusive for a very long period of time.
- It was only in 1953 that James Watson and Francis Crick, based on the X-ray diffraction data produced by Maurice Wilkins and Rosalind Franklin, proposed a very simple but famous Double Helix model for the structure of DNA. One of the hallmarks of their proposition was base pairing between the two strands of polynucleotide chains. However, this proposition was also based on the observation of Erwin Chargaff that for a double stranded DNA, the ratios between Adenine and Thymine and Guanine and Cytosine are constant and equals one.
CHARGAFF’S RULES
- E. Chargaff and his colleagues (1949) at Columbia analysed the composition of DNA from various sources by quantitative chromatographic methods. Their observations are called Chargaff’s rules, which state:
- Regardless of the source, the purine and pyrimidine bases occur in equal amounts in a DNA molecule, i.e., A + G = T + C.
- The amount of adenine is equivalent to the amount of thymine (A/T = 1) and the amount of cytosine is equivalent to that of guanine (C/G=1). Thus there is base equivalence in DNA.
- The base ratio, , however, rarely equal to 1, and varies with different species from 0.4 to 1.9 (low in microorganisms and high in higher animals), but is constant for a species. Therefore, base ratio has been used to identify the DNA from a particular source.
The base pairing confers a very unique property to the polynucleotide chains. They are said to be complementary to each other, and therefore if the sequence of bases in one strand is known then the sequence in other strand can be predicted. Also, if each strand from a DNA (let us call it as a parental DNA) acts as a template for synthesis of a new strand, the two double stranded DNA (let us call them as daughter DNA) thus, produced would be identical to the parental DNA molecule. Because of this, the genetic implications of the structure of DNA became very clear.
The salient features of the Double-helix structure of DNA are as follows:
- It is made of two polynucleotide chains, where the backbone is constituted by sugar-phosphate, and the bases project inside.
- The two chains have anti-parallel polarity. It means, if one chain has the polarity 5'3', the other has 3' 5' .
- The bases in two strands are paired through hydrogen bond (H-bonds) forming base pairs (bp). Adenine forms two hydrogen bonds with Thymine from opposite strand and vice-versa. Similarly, Guanine is bonded with Cytosine with three H-bonds. As a result, always a purine comes opposite to a pyrimidine. This generates approximately uniform distance between the two strands of the helix.
- The two chains are coiled in a right-handed fashion. The pitch of the helix is 3.4 nm (a nanometre is one billionth of a metre, that is 10–9 m) and there are roughly 10 bp in each turn. Consequently, the distance between a bp in a helix is approximately equal to 0.34 nm.
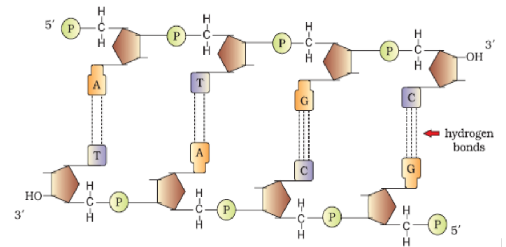
- The plane of one base pair stacks over the other in double helix. This, in addition to H-bonds, confers stability of the helical structure
Forms of DNA
- The double stranded DNA has five forms, A–, B–, C–, D– and Z–DNA. The first four forms (A–, B–, C– and D–DNA) are right handed while the last form (Z–DNA) is left handed. The B– form is the metabolically stable configuration and it is the structure proposed by Watson and Crick. In Z–DNA, phospho-diester backbone of the strands follows a zig-zag course, hence the name Z-DNA. It was discovered by Alexander Rich et. al. (1979). It is formed only when purines and pyrimidines are present alternately in the chain (or ). It is present in the salivary gland polytene chromosomes of Drosophila. It has a regulatory function.
Table: Comparison between various forms of DNA
| Chapters |
FORMS OF DNA |
||||
|
A–DNA |
B–DNA |
C–DNA |
D–DNA |
Z–DNA |
|
|
Direction of coiling |
Right handed |
Right handed |
Right handed |
Right handed |
Left handed |
|
Base pair per turn |
11 |
10 |
9 |
8 |
12 |
|
S-P Backbone |
Smooth |
Smooth |
Smooth |
Smooth |
Zig-Zag |
|
Axial rise (h) |
2.56Å |
3.37Å |
3.32Å |
3.03Å |
3.8Å |
|
Length of one pitch |
28.16Å |
34Å |
31Å |
24.24Å |
45.6Å |
|
Diameter |
23Å |
20Å |
19Å |
– |
18Å |
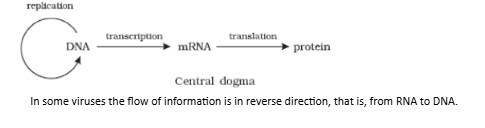
- The proposition of a double helix structure for DNA and its simplicity in explaining the genetic implication became revolutionary. Very soon, Francis Crick proposed the Central dogma in molecular biology, which states that the genetic information flows from DNARNAProtein.
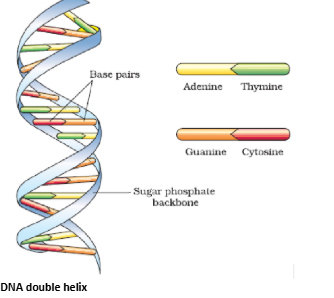
DNA Double Helix
Packaging of DNA Helix
- Taken the distance between two consecutive base pairs as 0.34 nm (0.34×10–9 m), if the length of DNA double helix in a typical mammalian cell is calculated (simply by multiplying the total number of bp with distance between two consecutive bp, that is, 6.6 × 109 bp × 0.34 × 10–9 m/bp), it comes out to be approximately 2.2 metres. A length that is far greater than the dimension of a typical nucleus (approximately 10–6 m).
- In prokaryotes, such as, E. coli, though they do not have a defined nucleus, the DNA is not scattered throughout the cell. DNA (being negatively charged) is held with some proteins (that have positive charges) in a region termed as ‘nucleoid’. The DNA in nucleoid is organised in large loops held by proteins.
- In eukaryotes, this organisation is much more complex. There is a set of positively charged, basic proteins called histones. A protein acquires charge depending upon the abundance of amino acids residues with charged side chains.
- Histones are rich in the basic amino acid residues lysines and arginines. Both the amino acid residues carry positive charges in their side chains. Histones are organised to form a unit of eight molecules called as histone octamer.
- The negatively charged DNA is wrapped around the positively charged histone octamer to form a structure called nucleosome. A typical nucleosome contains 200 bp of DNA helix. Nucleosomes constitute the repeating unit of a structure in nucleus called chromatin, thread-like stained (coloured) bodies seen in nucleus. The nucleosomes in chromatin are seen as ‘beads-on-string’ structure when viewed under electron microscope (EM).
- The beads-on-string structure in chromatin is packaged to form chromatin fibers that are further coiled and condensed at metaphase stage of cell division to form chromosomes. The packaging of chromatin at higher level requires additional set of proteins that collectively are referred to as
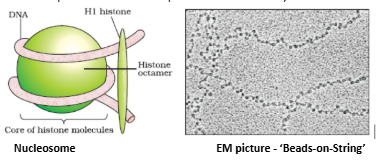
Non-histone Chromosomal (NHC) proteins
- In a typical nucleus, some region of chromatin are loosely packed (and stains light) and are referred to as euchromatin. The chromatin that is more densely packed and stains dark are called as Heterochromatin. Euchromatin is said to be transcriptionally active chromatin, whereas heterochromatin is inactive.
THE Search For Genetic Material
- Even though the discovery of nuclein by Meischer and the proposition for principles of inheritance by Mendel were almost at the same time, but that the DNA acts as a genetic material took long to be discovered and proven.
- By 1926, the quest to determine the mechanism for genetic inheritance had reached the molecular level. Previous discoveries by Gregor Mendel, Walter Sutton, Thomas Hunt Morgan and numerous other scientists had narrowed the search to the chromosomes located in the nucleus of most cells. But the question of what molecule was actually the genetic material, had not been answered.
Transforming Principle
- In 1928, Frederick Griffith, in a series of experiments with Streptococcus pneumoniae(bacterium responsible for pneumonia), witnessed a miraculous transformation in the bacteria. During the course of his experiment, a living organism (bacteria) had changed in physical form.
- When Streptococcus pneumoniae (pneumococcus) bacteria are grown on a culture plate, some produce smooth shiny colonies (S) while others produce rough colonies (R). This is because the S strain bacteria have a mucous (polysaccharide) coat, while R strain does not. Mice infected with the S strain (virulent) die from pneumonia infection but mice infected with the R strain do not develop pneumonia.

- Griffith was able to kill bacteria by heating them. He observed that heat-killed S strain bacteria injected into mice did not kill them. When he

injected a mixture of heat-killed S and live R bacteria, the mice died. Moreover, he recovered living S bacteria from the dead mice. He concluded that the R strain bacteria had somehow been transformed by the heat-killed S strain bacteria. Some ‘transforming principle’, transferred from the heat-killed S strain, had enabled the R strain to synthesise a smooth polysaccharide coat and become virulent. This must be due to the transfer of the genetic material. However, the biochemical nature of genetic material was not defined from his experiments.
Biochemical Characterisation of Transforming Principle
- Prior to the work of Oswald Avery, Colin MacLeod and Maclyn McCarty (1933-44), the genetic material was thought to be a protein. They worked to determine the biochemical nature of ‘transforming principle’ in Griffith's experiment.
- They purified biochemicals (proteins, DNA, RNA, etc.) from the heat-killed S cells to see which ones could transform live R cells into S cells. They discovered that DNA alone from S bacteria caused R bacteria to become transformed.
- They also discovered that protein-digesting enzymes (proteases) and RNA-digesting enzymes (RNases) did not affect transformation, so the transforming substance was not a protein or RNA. Digestion with DNase did inhibit transformation, suggesting that the DNA caused the transformation. They concluded that DNA is the hereditary material, but not all biologists were convinced.
The Genetic Material is DNA
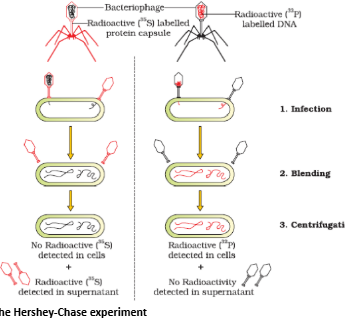
- The unequivocal proof that DNA is the genetic material came from the experiments of Alfred Hershey and Martha Chase (1952). They worked with viruses that infect bacteria called bacteriophages.
- The bacteriophage attaches to the bacteria and its genetic material then enters the bacterial cell. The bacterial cell treats the viral genetic material as if it was its own and subsequently manufactures more virus particles. Hershey and Chase worked to discover whether it was protein or DNA from the viruses that entered the bacteria.
- They grew some viruses on a medium that contained radioactive phosphorus and some others on medium that contained radioactive sulfur. Viruses grown in the presence of radioactive phosphorus contained radioactive DNA but not radioactive protein because DNA contains phosphorus but protein does not. Similarly, viruses grown on radioactive sulfur contained radioactive protein but not radioactive DNA because DNA does not contain sulfur.
- Radioactive phages were allowed to attach to E. coli bacteria. Then, as the infection proceeded, the viral coats were removed from the bacteria by agitating them in a blender. The virus particles were separated from the bacteria by spinning them in a centrifuge.
- Bacteria which was infected with viruses that had radioactive DNA were radioactive, indicating that DNA was the material that passed from the virus to the bacteria. Bacteria that were infected with viruses that had radioactive proteins were not radioactive. This indicates that proteins did not enter the bacteria from the viruses. DNA is therefore the genetic material that is passed from virus to bacteria.
Properties of Genetic Material (DNA versus RNA)
- It is clear that the debate between proteins versus DNA as the genetic material was unequivocally resolved from Hershey-Chase experiment. It became an established fact that it is DNA that acts as genetic material. However, it subsequently became clear that in some viruses, RNA is the genetic material (for example, Tobacco Mosaic viruses, QB bacteriophage, etc.).
- Answer to some of the questions such as, why DNA is the predominant genetic material, whereas RNA performs dynamic functions of messenger and adapter has to be found from the differences between chemical structures of the two nucleic acid molecules.
A molecule that can act as a genetic material must fulfill the following criteria:
- It should be able to generate its replica (Replication).
- It should chemically and structurally be stable.
- It should provide the scope for slow changes (mutation) that are required for evolution.
- It should be able to express itself in the form of 'Mendelian Characters’.
- If one examines each requirement one by one, because of rule of base pairing and complementarity, both the nucleic acids (DNA and RNA) have the ability to direct their duplications. The other molecules in the living system, such as proteins fail to fulfill first criteria itself.
- The genetic material should be stable enough not to change with different stages of life cycle, age or with change in physiology of the organism. Stability as one of the properties of genetic material was very evident in Griffith’s ‘transforming principle’ itself that heat, which killed the bacteria, at least did not destroy some of the properties of genetic material. This now can easily be explained in light of the DNA that the two strands being complementary if separated by heating come together, when appropriate conditions are provided. Further, 2'-OH group present at every nucleotide in RNA is a reactive group and makes RNA labile and easily degradable. RNA is also now known to be catalytic, hence reactive. Therefore, DNA chemically is less reactive and structurally more stable when compared to RNA. Therefore, among the two nucleic acids, the DNA is a better genetic material.
- In fact, the presence of thymine at the place of uracil also confers additional stability to DNA.
- Both DNA and RNA are able to mutate. In fact, RNA being unstable, mutate at a faster rate. Consequently, viruses having RNA genome and having shorter life span mutate and evolve faster. RNA can directly code for the synthesis of proteins, hence can easily express the characters. DNA, however, is dependent on RNA for synthesis of proteins. The protein synthesising machinery has evolved around RNA.
- The above discussion indicate that both RNA and DNA can function asgenetic material, but DNA being more stable is preferred for storage of genetic information. For the transmission of genetic information, RNA is better.
RNA World
- RNA was the first genetic material. There is now enough evidence to suggest that essential life processes (such as metabolism, translation, splicing, etc.), evolved around RNA. RNA used to act as a genetic material as well as a catalyst (there are some important biochemical reactions in living systems that are catalysed by RNA catalysts and not by protein enzymes). But, RNA being a catalyst was reactive and hence unstable. Therefore, DNA has evolved from RNA with chemical modifications that make it more stable. DNA being double stranded and having complementary strand further resists changes by evolving a process of repair.
Replication
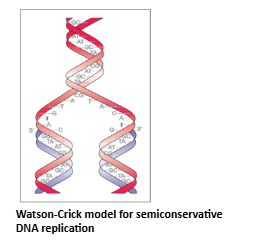
- While proposing the double helical structure for DNA, Watson and Crick had immediately proposed a scheme for replication of DNA. To quote their original statement that is as follows: ‘‘It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material’’ (Watson and Crick, 1953). The scheme suggested that the two strands would separate and act as a template for the synthesis of new complementary strands. After the completion of replication, each DNA molecule would have one parental and one newly synthesised strand. This scheme was termed as semiconservative DNA replication.
The Experimental Proof
- It is now proven that DNA replicates semiconservatively. It was shown first in Escherichia coli and subsequently in higher organisms, such as plants and human cells. Matthew Meselson and Franklin Stahl performed the following experiment in 1958:
They grew E. coli in a medium containing 15NH4Cl (15N is the heavy isotope of nitrogen) as the only nitrogen source for many generations. The result was that 15N was incorporated into newly synthesised DNA (as well as other nitrogen containing compounds). This heavy DNA molecule could be distinguished from the normal DNA by centrifugation in a cesium chloride (CsCl) density gradient (Note that 15N is not a radioactive isotope, and it can be separated from 14N only based on densities).
Then they transferred the cells into a medium with normal 14NH4Cl and took samples at various definite time intervals as the cells multiplied, and extracted the DNA that remained as double-stranded helices. The various samples were separated independently on CsCl gradients to measure the densities of DNA. The results are shown figure.
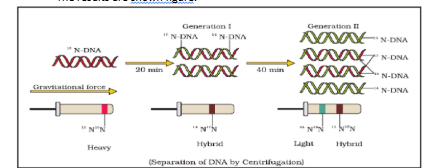
Thus, the DNA that was extracted from the culture one generation after the transfer from 15N to 14N medium [that is after 20 minutes; E. coli divides in 20 minutes] had a hybrid or intermediate density. DNA extracted from the culture after another generation [that is after 40 minutes, II generation] wascomposed of equal amounts of this hybrid DNA and of ‘light’ DNA.
- Very similar experiments involving use of radioactive thymidine to detect distribution of newly synthesised DNA in the chromosomes was performed on Vicia faba (faba beans) by Taylor and colleagues in 1958. The experiments proved that the DNA in chromosomes also replicate semiconservatively.
The Machinery and the Enzymes
- In living cells, such as E. coli, the process of replication requires a set of catalysts (enzymes). The main enzyme is referred to as DNA-dependent DNA polymerase, since it uses a DNA template to catalyse the polymerisation of deoxynucleotides.
- These enzymes are highly efficient enzymes as they have to catalyse polymerisation of a large number of nucleotides in a very short time. E. coli that has only 4.6 ×106 bp (compare it with human whose diploid content is 6.6 × 109 bp), completes the process of replication within 38 minutes; that means the average rate of polymerisation has to be approximately 2000 bp per second.
- Not only do these polymerases have to be fast, but they also have to catalyse the reaction with high degree of accuracy. Any mistake during replication would result into mutations. Furthermore, energetically replication is a very expensive process. Deoxyribonucleoside triphosphates serve dual purposes. In addition to acting as substrates, they provide energy for polymerization reaction (the two terminal phosphates in a deoxynucleoside triphosphates are high-energy phosphates, same as in case of ATP).
- In addition to DNA-dependent DNA polymerases, many additional enzymes are required to complete the process of replication with high degree of accuracy.
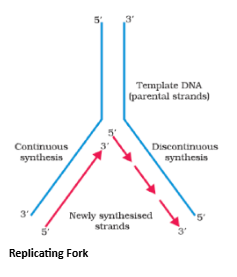
- For long DNA molecules, since the two strands of DNA cannot be separated in its entire length (due to very high energy requirement), the replication occur within a small opening of the DNA helix, referred to as replication fork.
- The DNA-dependent DNA polymerases catalyse polymerisation only in one direction, that is 5'3'. This creates some additional complications at the replicating fork. Consequently, on one strand (the template with polarity 3'5' ), the replication is continuous, while on the other (the template with polarity 5'3' ), it is discontinuous. The discontinuously synthesized fragments are later joined by the enzyme DNA ligase.
- The DNA polymerases on their own cannot initiate the process of replication. Also the replication does not initiate randomly at any place in DNA. There is a definite region in E. coli DNA where the replication originates. Such regions are termed as origin of replication.
- It isbecause of the requirement of the origin of replication that a piece of DNA if needed to be propagated during recombinant DNA procedures, requires a vector. The vectors provide the origin of replication.
- In eukaryotes, the replication of DNA takes place at S-phase of the cell-cycle. The replication of DNA and cell division cycle should be highly coordinated. A failure in cell division after DNA replication results into polyploidy (a chromosomal anomaly).
Mechanism of DNA Replication
- DNA replication is a multi-step complex process, which requires over a dozen enzymes, protein factors and metal ions. DNA replication takes place through following steps:
Initiation
- Replication begins at a particular spot called origin of replication (ori). Bacterial and viral DNA has a single origin of replication but eukaryotic DNA has a number of origins of replication because of the larger size as well as association with proteins. As a result, the whole prokaryotic DNA acts as a single replicating unit or replicon and eukaryotic DNA molecule has several replicating segments or replicons, each with its own origin and
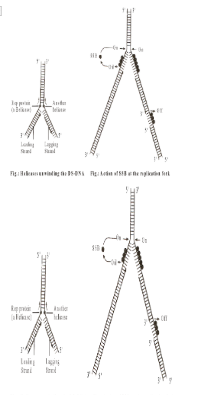
Terminus (where the origin of the next starts). In the circular DNA of prokaryotes the points of origin of replication and termination of replication are the same (Cairns, 1953). DNA replication can be unidirectional or bi-directional depending upon whether the replication from the point of origin proceeds only in one direction or proceeds in both the direction.
- Initiation of replication takes place due to a multi-enzyme complex called replisome or replication apparatus.
Activation of Deoxyribonucleotides
- All the deoxyribonucleoside monophosphates– dAMP (deoxyadenosine monophosphate), dGMP (deoxyguanosine monophosphate), dCMP (deoxycytidine monophosphate) and dTMP (deoxythymidine monophosphate) found floating free in the nucleoplasm serve as the raw materials in DNA synthesis. In the presence of energy (ATP) and an enzyme phosphorylase, the nucleotides dAMP, dGMP, dCMP and dTMP are phosphorylated and changed into triphosphate states dATP, dGTP, dCTP and dTTP, respectively.
Unwinding of parental DNA
- The DNA molecule is intricately coiled in its chromosome and its unwinding is not an easy process.
- The replication starts with an incision or a cut generally referred to as nick at a specific point in one of the two parental strands. The nick is brought about by an incision enzyme topoisomerase. As a result, origin of replication is identified. In the presence of enzyme helicases (unwindases) and a special protein (DNA unwinding protein), the two strands of DNA helix separate. In prokaryotes topoisomerases and helicases are replaced by DNA gyrases. The enzyme topoisomerase may seal the nick. The separated strands are stabilized by helix destabilizing (HD) or DNA binding proteins. Now they become open for replication and act as templates. However, whole of DNA does not open in one stretch but the point of separation proceeds slowly from one end to the other. It gives the appearance of Y-shaped structure called replication fork Replication initiates at end. Unwinding also creates a coiling tension ahead of the moving replication fork. This tension is reduced by topoisomerases.
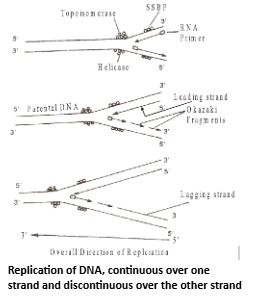
Base Pairing
- Deoxyribonucleoside triphosphates come to lie opposite the nitrogen bases of unwinded or exposed DNA template, base pairing being A= T and , i.e., dTTP opposite A, dCTP opposite G, dATP opposite T and dGTP opposite C.
- With the help of enzyme pyrophosphatase the two phosphates from the phosphorylated deoxyribonucleosides removed and change them into monophosphate state. Energy is released in this process.
Deoxyribouncleotide triphosphate Deoxyribonucleoside monophosphate + PP + Energy
dATPdAMP + 2Pi + energy
dGTP dGMP + 2Pi + energy
dCTPdCMP + 2Pi + energy
dTTPdTMP + 2Pi + energy
- The energy is used in establishing hydrogen bonds between the free nucleotides and nitrogen bases of template.
Chain formation
- In the presence of enzyme DNA polymerase III in prokaryotes and DNA polymerases and in eukaryotes (Kornberg, 1956), Mg++, ATP, GTP, TTP the adjacent nucleotides present attached to nitrogen bases of each template DNA strand establish phosphodiester bonds and get linked to form replicated DNA strand. As replication proceeds, new areas of parent DNA duplex unwind and separate so that replication proceeds rapidly from the place of origin towards the other ends. RNA primer is removed and the gap is filled with complementary nucleotides by means of DNA polymerase I. Because of sequential opening of DNA chain and its replication to form two chains, DNA replication is also called zipper duplication.
- However, DNA polymerase can polymerize nucleotides only in direction because it adds them at the end. Since the two strands of DNA run in antiparallel directions, the two templates provide different ends for replication. Replication occurs over the two templates thus proceeds in opposite directions. One strand is formed continuously because its end is open for elongation. It is called leading strand. Replication is not continuous on the second template because only a short segment of DNA strand can be build in direction due to exposure of a small stretch of template at one time.
- Short segments of replicated DNA are called Okazaki fragments(Reiji Okazaki, 1968). Each of them has 3000–4000 nucleotides. An RNA primer is also required every time as a new Okazaki fragment is to be built. After replacing RNA primer with deoxyribonucleotides and their polymerization, Okazaki fragments are joined together by means of enzyme, DNA ligase. DNA strand build up of Okazaki fragment is called lagging strand.
- Thus DNA replication is said to the semi-discontinuous because the leading strand is synthesized continuously and the lagging strand is formed discontinuously in short pieces that later join.
Table: Differences between leading and lagging strands of replicating DNA
|
LEADING STRAND |
Lagging strand |
|
1. It is a replicated strand of DNA, which grows continuously without any gap. 2. It needs a single RNA primer to start its growth. 3. It does not require DNA ligase for its growth. 4. The direction of growth of the leading strand is . |
1. Lagging strand is a replicated strand of DNA, which is, formed in short segments called Okazaki fragments. 2. Each segment needs a separate RNA primer to start. 3. DNA ligase is required for joining Okazaki segment. 4. The direction of growth of the lagging strand is though each Okazaki fragment grows in direction. |
Proof Reading (Editing)
- The specificity of base pairing ensures accurate replication. However, a wrong base is sometimes introduced during replication. DNA polymerase is able to sense the same. However, even DNA polymerase III is unable to distinguish uracil from thymine so that it is often incorporated in place of thymine. A nick or break is caused by an endonuclease near the region of repair. DNA polymerase III removes the mismatched or wrong nucleotides if present and DNA polymerase I synthesizes a correct replacement by using the intact strand as template. The newly formed segment is sealed by DNA ligase. Each daughter double DNA molecule becomes spirally coiled to form a double helix.
Transcription
- The process of copying genetic information from one strand of the DNA into RNA is termed as transcription. Here also, the principle of complementarity governs the process of transcription, except the adenosine now forms base pair with uracil instead of thymine. However, unlike in the process of replication, which once set in, the total DNA of an organism gets duplicated, in transcription only a segment of DNA and only one of the strands is copied into RNA. This necessitates defining the boundaries that would demarcate the region and the strand of DNA that would be transcribed.
- Why both the strands are not copied during transcription ?
- First, if both strands act a template, they would code for RNA molecule with different sequences (Remember complementarity does not mean identical), and in turn, if they code for proteins, the sequence of amino acids in the proteins would be different. Hence, one segment of the DNA would be coding for two different proteins, and this would complicate the genetic information transfer machinery.
- Second, the two RNA molecules if produced simultaneously would be complementary to each other, hence would form a double stranded RNA. This would prevent RNA from being translated into protein and the exercise of transcription would become a futile one.
Transcription Unit
- A transcription unit in DNA is defined primarily by the three regions in the DNA:
(i) A Promoter(ii) The Structural gene(iii) A Terminator
- There is a convention in defining the two strands of the DNA in the structural gene of a transcription unit. Since the two strands have opposite polarity and the DNA-dependent RNA polymerase also catalyse the polymerisation in only one direction, that is, 5'→3' , the strand that has the polarity 3'→5' acts as a template, and is also referred to as template strand.
- The other strand which has the polarity (5'→3') and the sequence same as RNA (except thymine at the place of uracil), is displaced during transcription. Strangely, this strand (which does not code for anything) is referred to as coding strand. All the reference point while defining a transcription unit is made with coding strand. To explain the point, a hypothetical sequence from a transcription unit is represented below:
3'-ATGCATGCATGCATGCATGCATGC-5' Template Strand
5'-TACGTACGTACGTACGTACGTACG-3' Coding Strand
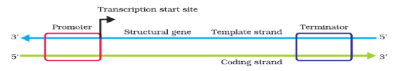
Schematic Structure of a Transcription Unit
- The promoter and terminator flank the structural gene in a transcription unit. The promoter is said to be located towards 5'-end (upstream) of the structural gene (the reference is made with respect to the polarity of coding strand). It is a DNA sequence that provides binding site for RNA polymerase, and it is the presence of a promoter in a transcription unit that also defines the template and coding strands. By switching its position with terminator, the definition of coding and template strands could be reversed. The terminator is located towards 3'-end (downstream) of the coding strand and it usually defines the end of the process of transcription. There are additional regulatory sequences that may be present further upstream or downstream to the promoter.
Transcription Unit and the Gene
- A gene is defined as the functional unit of inheritance. Though there is no ambiguity that the genes are located on the DNA, it is difficult to literally define a gene in terms of DNA sequence. The DNA sequence coding for tRNA or rRNA molecule also define a gene. However by defining a cistron as a segment of DNA coding for a polypeptide, the structural gene in a transcription unit could be said as monocistronic (mostly in eukaryotes) or polycistronic (mostly in bacteria or prokaryotes).
- In eukaryotes, the monocistronic structural genes have interrupted coding sequences-the genes in eukaryotes are split. The coding sequences or expressed sequences are defined as exons. Exons are said to be those sequence that appear in mature or processed RNA. The exons are interrupted by introns. Introns or intervening sequences do not appear in mature or processed RNA. The split-gene arrangement further complicates the definition of a gene in terms of a DNA segment. Inheritance of a character is also affected by promoter and regulatory sequences of a structural gene. Hence, sometime the regulatory sequences are loosely defined as regulatory genes, even though these sequences do not code for any RNA or protein.
Types of RNA and the process of Transcription
- In bacteria, there are three major types of RNAs: mRNA (messenger RNA), tRNA (transfer RNA), and rRNA (ribosomal RNA). All three RNAs are needed to synthesise a protein in a cell. The mRNA provides the template, tRNA brings aminoacids and reads the genetic code, and rRNAs play structural and catalytic role during translation.
- There is single DNA-dependent RNA polymerase that catalyses transcription of all types of RNA in bacteria. RNA polymerase binds to promoter and initiates transcription (Initiation). It uses nucleoside triphosphates as substrate
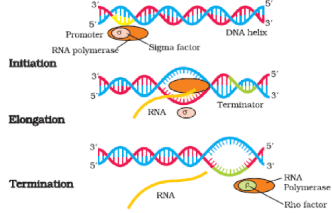
and polymerises in a template depended fashion following the rule of complementarity. It somehow also facilitates opening of the helix and continues elongation. Only a short stretch of RNA remains bound to the enzyme. Once the polymerases reaches the terminator region, the nascent RNA falls off, so also the RNA polymerase. This results in termination of transcription.
- An intriguing question is that how is the RNA polymerases able to catalyse all the three steps, which are initiation, elongation and termination. The RNA polymerase is only capable of catalysing the process of elongation. It associates transiently with initiation-factor (σ) and termination-factor (ρ) to initiate and terminate the transcription, respectively. Association with these factors alter the specificity of the RNA polymerase to either initiate or terminate.
- In bacteria, since the mRNA does not require any processing to become active, and also since transcription and translation take place in the same compartment (there is no separation of cytosol and nucleus in bacteria), many times the translation can begin much before the mRNA is fully transcribed. Consequently, the transcription and translation can be coupled in bacteria. In eukaryotes, there are two additional complexities:
(i)There are at least three RNA polymerases in the nucleus (in addition to the RNA polymerase found in the organelles). There is a clear cut division of labour. The RNA polymerase I transcribes rRNAs
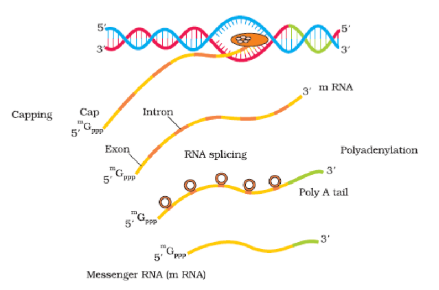
Process of Transcription in Eukaryotes
(28S, 18S, and 5.8S), whereas the RNA polymerase III is responsible for transcription of tRNA, 5srRNA, and snRNAs (small nuclear RNAs). The RNA polymerase II transcribes precursor of mRNA, the heterogeneous nuclear RNA (hnRNA).
The second complexity is that the primary transcripts contain both the exons and the introns and are non-functional. Hence, it is subjected to a process called splicing where the introns are removed and exons are joined in a defined order. hnRNA undergo two additional processing called as capping and tailing. In capping an unusual nucleotide (methyl guanosine triphosphate) is added to the 5'-end of hnRNA. In tailing, adenylate residues (200-300) are added at 3'-end in a template independent manner. It is the fully processed hnRNA, now called mRNA, that is transported out of the nucleus for translation.
- The significance of such complexities is now beginning to be understood. The split-gene arrangements represent probably an ancient feature of the genome. The presence of introns is reminiscent of antiquity, and the process of splicing represents the dominance of RNA-world. In recent times, the understanding of RNA and RNA-dependent processes in the living system have assumed more importance.
- Transcribing segment i.e., cistron has a promoter region where initiation of transcription begins, and a terminator region where transcription ends. Promoter has different parts for attachment to various transcription factors. In many cases, the promoter has an AT rich sequence called TATA box. The area has a groove to which specific protein components can combine. TATA containing region is also called Pribnow box after the name of its discoverer.
- Enzyme required for transcription is RNA polymerase (Chemberlain and Berg, 1962; isolated from E.coli). Eukaryotic RNA polymerases also require transcription factors for initiation. The entire enzyme (holozyme) consists of a core enzyme and a sigma factor. The core enzyme consists of four polypeptide chains, , and . The holozyme has a molecular weight . RNA polymerase not only initiates but also extends the RNA (chain elongation) and functions always in and direction.
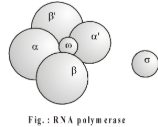
- Sigma factor recognizes the start signal or promoter region of DNA.
- Prokaryotes have only one type of RNA polymerase, which synthesizes all types of RNA. Eukaryotes have three types of RNA polymerases, I for rRNA, II for mRNA and III for tRNAs. A specific protein called rho factor (PR) is required for termination of transcription.
- In many bacteria, genes of related functions are grouped together in operons. An operon acts as a single transcription unit and thus produces polycistronic mRNA. In eukaryotes, only monocistronic mRNAs are generally produced.
- Actinomycin D prevents transcription.
Genetic Code
- During replication and transcription a nucleic acid was copied to form another nucleic acid. Hence, these processes are easy to conceptualise on the basis of complementarity. The process of translation requires
- Transfer of genetic information from a polymer of nucleotides to a polymer of amino acids. Neither does any complementarity exist between nucleotides and amino acids, nor could any be drawn theoretically. There existed ample evidences, though, to support the notion that change in nucleic acids (genetic material) were responsible for change in amino acids in proteins. This led to the proposition of a genetic code that could direct the sequence of amino acids during synthesis of proteins. If determining the biochemical nature of genetic material and the structure of DNA was very exciting, the proposition and deciphering of genetic code were most challenging. In a very true sense, it required involvement of scientists from several disciplines – physicists, organic chemists, biochemists and geneticists.
- It was George Gamow, a physicist, who argued that since there are only 4 bases and if they have to code for 20 amino acids, the code should constitute a combination of bases. He suggested that in order to code for all the 20 amino acids, the code should be made up of three nucleotides. This was a very bold proposition, because a permutation combination of 43 (4 × 4 × 4) would generate 64 codons; generating many more codons than required. Providing proof that the codon was a triplet, was a more daunting task.
- The chemical method developed by Har Gobind Khorana was instrumental in synthesising RNA molecules with defined combinations of bases (homopolymers and copolymers). Marshall Nirenberg’s cell-free system for protein synthesis finally helped the code to be deciphered. Severo Ochoa enzyme (polynucleotide phosphorylase) was also helpful in polymerising RNA with defined sequences in a template independent manner (enzymatic synthesis of RNA). Finally a checker-board for genetic code was prepared which is given in table.
Properties of Genetic Code
Non overlapping code: Studies on gene mutations in TMV show that the genetic code is non-overlapping, i.e., a nitrogenous base is a constituent of only one codon.
Commaless code: There is no gap or a base between two codons, i.e., it is continuous and having no punctuation marks (gaps) between the coding triplets.
Polarity: The genetic code is read in a fixed direction i.e., the code has polarity.

Having complementary anticodons: The genetic code has complementary anticodon, i.e.,
- CodonAUG
- AnticodonUAC
Start signals (Initial codons): Chain initiation codon is generally AUG, which codes methionine (in eukaryotes) or N-formyl methionine (in prokaryotes). Rarely GUG, which normally codes for valine, serves as the initiation codon in bacterial protein synthesis. Thus, both AUG and GUG codons show ambiguity.
Stop signals (Termination codons): Three of 64 codons do not code for any amino acids and hence cause termination and release of polypeptide chains. These codons are UAG (amber). UAA (ochre) and UGA (opal or umber). They are called termination codons or stop codons but previously nonsense codon. Termination codons are not read by tRNA molecule but by specific proteins called release factor (RF)- three in prokaryotes and single in eukaryotes.
Degeneracy of codons: There are many more codons than amino acid types. When a particular amino acid has more than one codon, the codon in called degenerate. Only tryptophan (UGG) and methionine (AUG) are specified by single codon and all other amino acids are specified by 2–6 codons. Thus, the genetic code is degenerate, i.e., it lacks specificity and one amino acid often has more than one code triplet. In degenerate codons the first two nitrogen bases are similar while the third one is different. As the third nitrogen base has no effect on coding, the same is called wobble position. Matthaei and Nirenberg discovered degeneracy of genetic code.
Non-ambiguous codon: When a particular codon codes for more than one amino acid, the codon is called ambiguous. There is no ambiguity regarding genetic code, i.e., one condon codes for same specific amino acid. But the codons like GGA, UGA, etc., show ambiguity. GGA codes for both glycine and glutamic acid. In yeast mitochondria, UGA codes for tryptophan, however, in other cases, it is termination codon.
Wobble Hypothesis: It was proposed by Crick (1966) to describe the lack of specificity in the third base of codon. According to this hypothesis, only the first two positions of a triplet codon on mRNA have a precise pairing with the bases of the tRNA anticodon. The pairing of the third positon bases of the condon may be ambiguous, and varies according to the nucleotide present at this position. Thus, a single tRNA type is able to recognize two or more codons differing only in the third base. The anticodon UCG of serine tRNA recognizes two codons, AGC and AGU. The bonding between UCG and AGC follows the usual Watson-Crick paring pattern. In UCG–AGU paring, however, hydrogen bonding takes place between G and U, instead of normal pairing G=C. Such interaction between the third bases is referred to as wobble pairing.
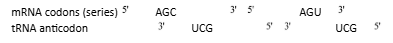
The wobbling allows economy of the number of tRNA molecules because several codons meant for same amino acid are recognized by same tRNA.
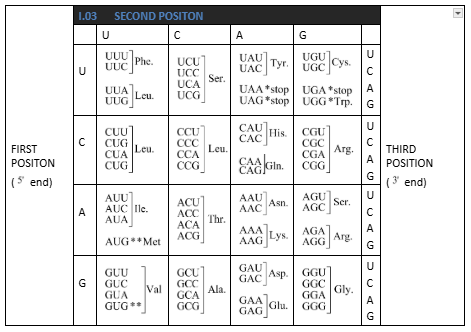
Table: The Codons for the Various Amino Acids
Mutations and Genetic Code
- The relationships between genes and DNA are best understood by mutation studies. Effects of large deletions and rearrangements in a segment of DNA are easy to comprehend. It may result in loss or gain of a gene and so a function. A classical example of point mutation is a change of single base pair in the gene for beta globin chain that results in the change of amino acid residue glutamate to valine.
- It results into a diseased condition called as sickle cell anemia. Effect of point mutations that inserts or deletes a base in structural gene can be better understood by following simple example. Consider a statement that is made up of the following words each having three letters like genetic code.
RAM HAS RED CAP: If we insert a letter B in between HAS and RED and rearrange the statement, it would read as follows:
RAM HAS BRE DCA P: Similarly, if we now insert two letters at the same place, say BI'. Now it would read,
RAM HAS BIR EDC AP: Now we insert three letters together, say BIG, the statement would read
RAM HAS BIG RED CAP: The same exercise can be repeated, by deleting the letters R, E and D, one by one and rearranging the statement to make a triplet word.
- RAM HAS EDC AP
- RAM HAS DCA P
- RAM HAS CAP
The conclusion from the above exercise is very obvious. Insertion or deletion of one or two bases changes the reading frame from the point of insertion or deletion. Insertion or deletion of three or its multiple bases insert or delete one or multiple codon hence one or multiple amino acids, and reading frame remains unaltered from that point onwards. Such mutations are referred to as frame-shift insertion or deletion mutations. This forms the genetic basis of proof that codon is a triplet and it is read in a contiguous manner.
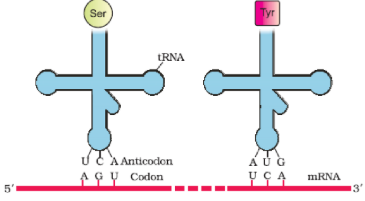
tRNA– the Adapter Molecule
- From the very beginning of the proposition of code, it was clear to Francis Crick that there has to be a mechanism to read the code and also to link it to the amino acids, because amino acids have no structural specialities to read the code uniquely. He postulated the presence of an adapter molecule that would on one hand read the code and on other hand would bind to specific amino acids. The tRNA, then called sRNA (soluble RNA), was known before the genetic code was postulated. However, its role as an adapter molecule was assigned much later. tRNA has an anticodon loop that has bases complementary to the code, and it also has an amino acid accepter end to which it binds to amino acids. tRNAs are specific for each amino acid.
For initiation, there is another specific tRNA that is referred to as initiator tRNA. There are no tRNAs for stop codons. The secondary structure of tRNA looks like a clover-leaf. In actual structure, the tRNA is a compact molecule which looks like inverted L.
Translation
- Translation refers to the process of polymerisation of amino acids to form a polypeptide. The order and sequence of amino acids are defined by the sequence of bases in the mRNA.
- The amino acids are joined by a bond which is known as a peptide bond. Formation of a peptide bond requires energy. Therefore, in the first phase itself amino acids are activated in the presence of ATP and linked to their cognate tRNA– a process commonly called as charging of tRNA or aminoacylation of tRNA to be more specific.
- If two such charged tRNAs are brought close enough, the formation of peptide bond between them would be favoured energetically. The presence of a catalyst would enhance the rate of peptide bond formation. The cellular factory responsible for synthesising proteins is the ribosome. The ribosome consists of structural RNAs and about 80 different proteins. In its inactive state, it exists as two subunits; a large subunit and a small subunit. When the small subunit encounters an mRNA, the process of translation of the mRNA to protein begins.
- There are two sites in the large subunit, for subsequent amino acids to bind to and thus, be close enough to each other for the formation of a peptide bond. The ribosome also acts as a catalyst (23S rRNA in bacteria is the enzyme- ribozyme) for the formation of peptide bond.
- A translational unit in mRNA is the sequence of RNA that is flanked by the start codon (AUG) and the stop codon and codes for a polypeptide. An mRNA also has some additional sequences that are not translated and are referred as untranslated regions (UTR). The UTRs are present at both 5' -end (before start codon) and at 3' -end (after stop codon). They are required for efficient translation process.
- For initiation, the ribosome binds to the mRNA at the start codon (AUG) that is recognised only by the initiator tRNA. The ribosome proceeds to the elongation phase of protein synthesis. During this stage, complexes composed of an amino acid linked to tRNA, sequentially bind to the appropriate codon in mRNA by forming complementary base pairs with the tRNA anticodon. The ribosome moves from codon to codon along the mRNA. Amino acids are added one by one, translated into Polypeptide sequences dictated by DNA and represented by mRNA. At the end, a release factor binds to the stop codon, terminating translation and releasing the complete polypeptide from the ribosome.
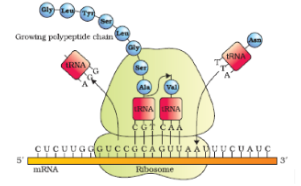
Translation (PROTEIN SYNTHESIS) - STEPS
- It is the process in which the coded genetic message brought by mRNA from DNA is changed into a polypeptide chain with a sequence of amino acids similar to the sequence of codons present over mRNA.
- This occurs in the cytoplasm in both eukaryotes (also in mitochondria and plastids) and prokaryotes. Besides mRNA, the three key players in this process are: ribosomes, tRNA and amino acids. The process of translation involves following steps.
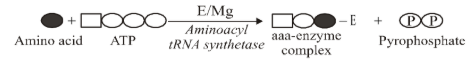
Activation of amino acids
- It is the first step in translation. Amino acids are activated with ATP in the presence of an activating enzyme, aminoacyl tRNA synthetase (Zamecnick and Hoagland, 1957) to from an “energy rich enzyme bond” amino acyl adenylate (aaa) enzyme complex. Mg++ is also required. Pyrophosphate (PPi) is released in this process.
Transfer of activated amino acids to tRNA (Charging of tRNA)
- Amino-acyl-adenylate enzyme complex colloids with free floating tRNA molecules. The amino acid joins at its carboxyl end (–COOH) with the ACC– () end of tRNA to form amino-acyl-tRNA complex (charged tRNA) or AA-tRNA. The enzyme (aminoacyl synthetase) and AMP are released. The transfer is catalysed by the same enzyme, aminoacyl synthetase. However, there is a specific enzyme for each kind of amino acid.
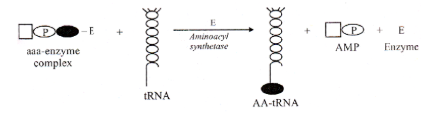
- The free enzyme can activate and attach another amino acid molecule to another tRNA molecule. The energy released by change of ATP to AMP is retained in the amino acid tRNA complex (charged tRNA). The energy is later used to drive the formation of peptide bond when amino acids link together on ribosomes. The charged tRNA moves to the site of protein synthesis, the ribosome.
Initiation of Polypeptide Chain
- The initiation of protein synthesis requires certain initiation factors (IF).
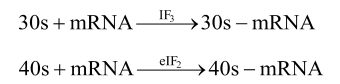
- There are three initiation factors in prokaryotes (IF3, IF2 and IF1) and nine in eukaryotes.
- The starting amino acid in bacterial polypeptide is N-formyl methionine and in eukaryotes is methionine.
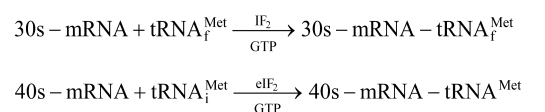
- The larger subunit of ribosome combines with smaller subunit –tRNAMet complex to form functional intact ribosome (70S or 80S).
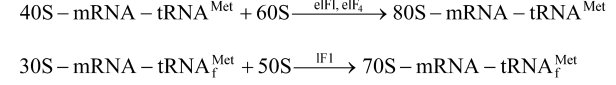
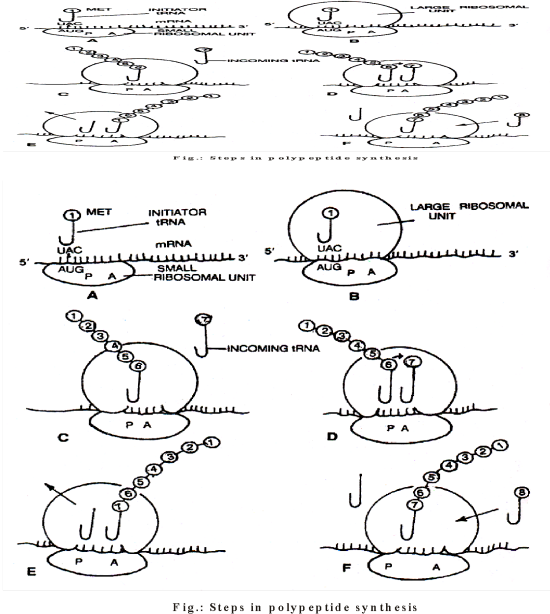
Elongation of Polypeptide Chain
- An aminoacyl tRNA complex reaches the A-site and attaches to mRNA codon next to initiation codon with the help of its anticodon.
- A peptide bond (–CO–NH–) is established between the carboxyl group of amino acid attached to tRNA at P-site and amino group of amino acid attached to tRNA at A-site.
- This is catalysed by the peptidyl transferase enzyme located in the large subunit.
- The process of translocation requires a factor called translocase (EF–G factor) in prokaryotes and eEF2 in eukaryotes, and energy from GTP.
Termination of Polypeptide Chain
- Polypeptide synthesis stops when a termination codon (UAA, UAG or UGA) reaches the A-site.
- The P-site tRNA is hydrolysed and the complete polypeptide is released in the presence of GTP dependent release factor (eRF1 in eukaryotes, and RF1 and RF2 in prokaryotes).
- The two subunits of ribosome also dissociate in the presence of dissociation factor (DF).
Modification of Released polypeptide
The just released polypeptide has primary structure. It may lose some amino acids from the end with the help of an exopeptidase enzyme, and then coil and fold on itself to acquire secondary and tertiary structures. Folding takes place with the help of special proteins called molecular chaperons.
Polysome formation
When the ribosome has moved sufficiently down the mRNA chain towards 3' end, another ribosome takes up position at the initiator codon of mRNA. Synthesis of many molecules of the same polypeptide simultaneously from one mRNA molecule by a polysome is called translational amplification.
Energy used
One GTP is hydrolysed as each successive amino acid–tRNA complex attached to the A-site. A second GTP is broken down as the ribosome moves to each new codon. One ATP is hydrolysed during amino acid activation. Thus the formation of each peptide bond uses 3 high-energy molecules, one ATP and 2GTP.
Antibiotics and Bacterial protein synthesis
| Antibiotic | Effect |
|---|---|
| Streptomycin | Inhibits initiation of translation and causes misreading. |
| Tetracycline | Inhibits binding of aminoacyl tRNA to ribosome |
| Neomycin | Inhibits interaction between tRNA and mRNA. |
| Chloramphenicol | Inhibits peptidyl transferase and so formation of peptide bonds. |
| Erythromycin | Inhibits translocation of mRNA along ribosome. |
Regulation of Gene Expression
Regulation of gene expression refers to a very broad term that may occur at various levels. Considering that gene expression results in the formation of a polypeptide, it can be regulated at several levels. In eukaryotes, the regulation could be exerted at:
- Transcriptional level (formation of primary transcript)
- Processing level (regulation of splicing)
- Transport of mRNA from nucleus to the cytoplasm
- Translational level
The genes in a cell are expressed to perform a particular function or a set of functions. For example, if an enzyme called beta-galactosidase is synthesised by E. coli, it is used to catalyse the hydrolysis of lactose into galactose and glucose. Therefore, in simple terms, it is the metabolic, physiological or environmental conditions that regulate the expression of genes.
The Lac operon
The elucidation of the lac operon was a result of a close association between a geneticist, Francois Jacob and a biochemist, Jacque Monod. They were the first to elucidate a transcriptionally regulated system.
In lac operon (here lac refers to lactose), a polycistronic structural gene is regulated by a common promoter and regulatory genes. Such arrangement is very common in bacteria and is referred to as operon.
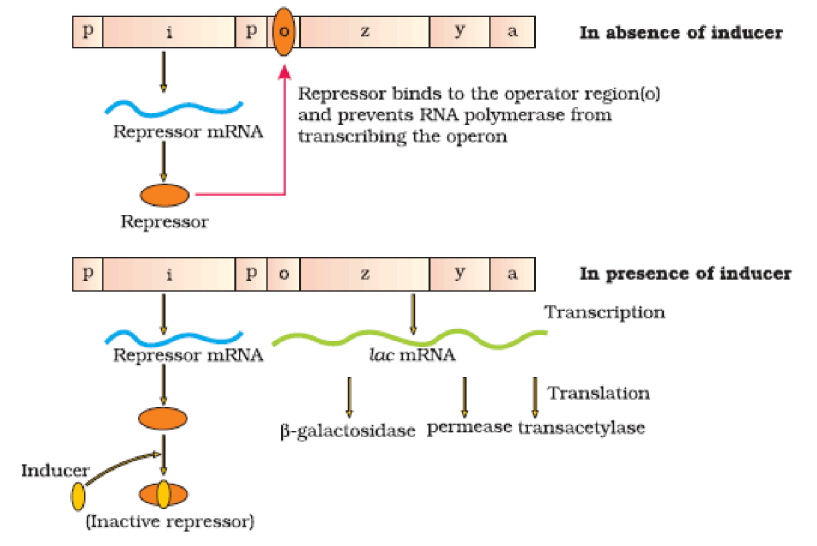
The lac operon consists of:
- One regulatory gene (i gene) - codes for the repressor of the lac operon
- Three structural genes (z, y, and a):
- z gene codes for beta-galactosidase (β-gal)
- y gene codes for permease
- a gene encodes a transacetylase
The lac Operon Mechanism
In absence of inducer:
- The repressor of the operon is synthesised (constitutively) from the i gene.
- The repressor protein binds to the operator region of the operon and prevents RNA polymerase from transcribing the operon.
In presence of inducer:
- Lactose is the substrate for the enzyme beta-galactosidase and it regulates switching on and off of the operon. Hence, it is termed as inducer.
- The repressor is inactivated by interaction with the inducer (lactose or allolactose).
- This allows RNA polymerase access to the promoter and transcription proceeds.
Regulation of lac operon by repressor is referred to as negative regulation. Lac operon is under control of positive regulation as well.
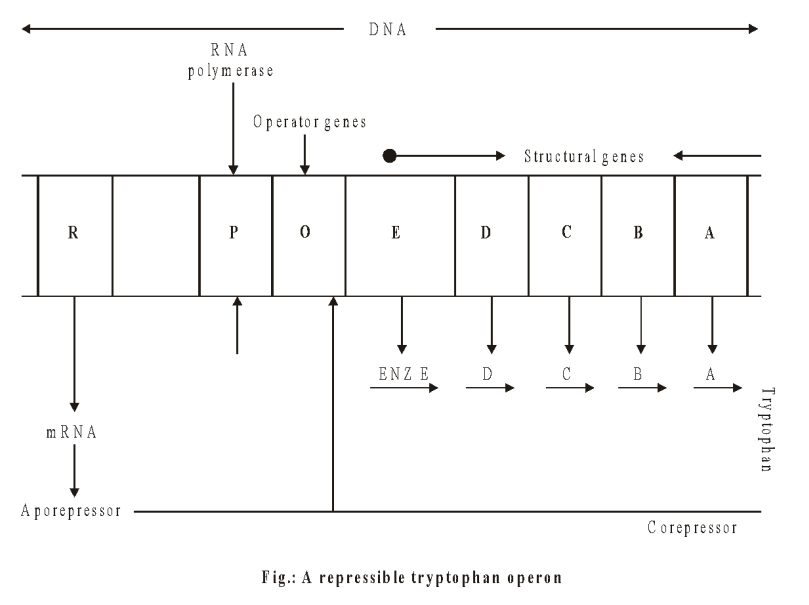
Repressible operon system/Tryptophan operon system
The repressible operon system is a regulated genetic material, which normally remains operational, and the enzymes produced by its structural genes are normally present in the cell. The functioning is stopped when the concentration of an end product crosses a threshold value or not required.
An example of repressible system is tryptophan or trp operon of E.coli. It consists of following components:
- Structural genes: trp E, trp D, trp C, trp B, trp A (code for five enzymes needed for tryptophan synthesis)
- Operator genes
- Promoter genes
- Regulator genes
- Aporepressor
- Corepressor
Mechanism:
- Operator gene is normally kept switched on because the aporepressor produced by regulator gene is unable to block operator gene.
- Aporepressor alone is unable to block the activity of operator gene because of the absence of the binding head.
- Corepressor (amino acid tryptophan) acts as a nonproteinous component of repressor.
- If tryptophan is available from outside, this binds with the aporepressor to produce repressor, which blocks the operator gene activity.
Human Genome Project
It is the sequence of bases in DNA that determines the genetic information of a given organism. If two individuals differ, then their DNA sequences should also be different, at least at some places. These assumptions led to the quest of finding out the complete DNA sequence of human genome.
Human Genome Project (HGP) was called a mega project and was launched in the year 1990. The project was completed in 2003.
- Human genome is said to have approximately 3 × 109 bp
- If the cost of sequencing required is US $ 3 per bp, the total estimated cost would be approximately 9 billion US dollars
- If stored in typed form in books (1000 letters per page, 1000 pages per book), then 3300 books would be required
- HGP was closely associated with the rapid development of Bioinformatics
Goals of HGP
- Identify all the approximately 20,000-25,000 genes in human DNA
- Determine the sequences of the 3 billion chemical base pairs that make up human DNA
- Store this information in databases
- Improve tools for data analysis
- Transfer related technologies to other sectors, such as industries
- Address the ethical, legal, and social issues (ELSI) that may arise from the project
The Human Genome Project was a 13-year project coordinated by the U.S. Department of Energy and the National Institute of Health. During the early years of the HGP, the Wellcome Trust (U.K.) became a major partner; additional contributions came from Japan, France, Germany, China and others.
Methodologies
The methods involved two major approaches:
Approach 1: Expressed Sequence Tags (ESTs): One approach focused on identifying all the genes that expressed as RNA (referred to as Expressed Sequence Tags).
Approach 2: Whole Genome Sequencing
The other took the blind approach of simply sequencing the whole set of genome that contained all the coding and non-coding sequence, and later assigning different regions in the sequence with functions (a term referred to as Sequence Annotation).
Sequencing Process:
- Total DNA from a cell is isolated and converted into random fragments of relatively smaller sizes
- Cloned in suitable host using specialised vectors
- The commonly used hosts were bacteria and yeast
- The vectors were called as BAC (bacterial artificial chromosomes), and YAC (yeast artificial chromosomes)
- The fragments were sequenced using automated DNA sequencers that worked on the principle of a method developed by Frederick Sanger
Salient Features of Human Genome
- The human genome contains 3164.7 million nucleotide bases.
- The average gene consists of 3000 bases, but sizes vary greatly, with the largest known human gene being dystrophin at 2.4 million bases.
- The total number of genes is estimated at 30,000–much lower than previous estimates of 80,000 to 1,40,000 genes.
- Almost all (99.9 per cent) nucleotide bases are exactly the same in all people.
- The functions are unknown for over 50 per cent of discovered genes.
- Less than 2 per cent of the genome codes for proteins.
- Repeated sequences make up very large portion of the human genome.
- Chromosome 1 has most genes (2968), and the Y has the fewest (231).
- Scientists have identified about 1.4 million locations where single base DNA differences (SNPs – single nucleotide polymorphism) occur in humans.
Applications and Future Challenges
- Deriving meaningful knowledge from the DNA sequences will define research through the coming decades
- One of the greatest impacts may well be enabling a radically new approach to biological research
- In the past, researchers studied one or a few genes at a time
- With whole-genome sequences, we can approach questions systematically and on a much broader scale
DNA Fingerpriting
99.9 per cent of base sequence among humans is the same. It is these differences in sequence of DNA which make every individual unique in their phenotypic appearance. DNA fingerprinting is a very quick way to compare the DNA sequences of any two individuals.
DNA fingerprinting involves identifying differences in some specific regions in DNA sequence called as repetitive DNA, because in these sequences, a small stretch of DNA is repeated many times. These repetitive DNA are separated from bulk genomic DNA as different peaks during density gradient centrifugation.
Satellite DNA: The bulk DNA forms a major peak and the other small peaks are referred to as satellite DNA. Depending on base composition, length of segment, and number of repetitive units, the satellite DNA is classified into many categories, such as:
- Micro-satellites
- Mini-satellites
Applications of DNA Fingerprinting
- Forensic applications: Since DNA from every tissue shows the same degree of polymorphism, they become very useful identification tool
- Paternity testing: As the polymorphisms are inheritable from parents to children
- Genetic mapping of human genome
- Population and genetic diversities
Polymorphism (variation at genetic level) arises due to mutations. If a germ cell mutation does not seriously impair individual's ability to have offspring, it can spread to other members of population through sexual reproduction.
DNA Fingerprinting Technique
The technique of DNA Fingerprinting was initially developed by Alec Jeffreys. He used a satellite DNA as probe that shows very high degree of polymorphism. It was called as Variable Number of Tandem Repeats (VNTR).
The technique included:
- Isolation of DNA
- Digestion of DNA by restriction endonucleases
- Separation of DNA fragments by electrophoresis
- Transferring (blotting) of separated DNA fragments to synthetic membranes, such as nitrocellulose or nylon
- Hybridisation using labelled VNTR probe
- Detection of hybridised DNA fragments by autoradiography
The VNTR belongs to a class of satellite DNA referred to as mini-satellite. A small DNA sequence is arranged tandemly in many copy numbers. The copy number varies from chromosome to chromosome in an individual. The numbers of repeat show very high degree of polymorphism. As a result the size of VNTR varies in size from 0.1 to 20 kb.
After hybridisation with VNTR probe, the autoradiogram gives many bands of differing sizes. These bands give a characteristic pattern for an individual DNA. It differs from individual to individual in a population except in the case of monozygotic (identical) twins. The sensitivity of the technique has been increased by use of polymerase chain reaction.
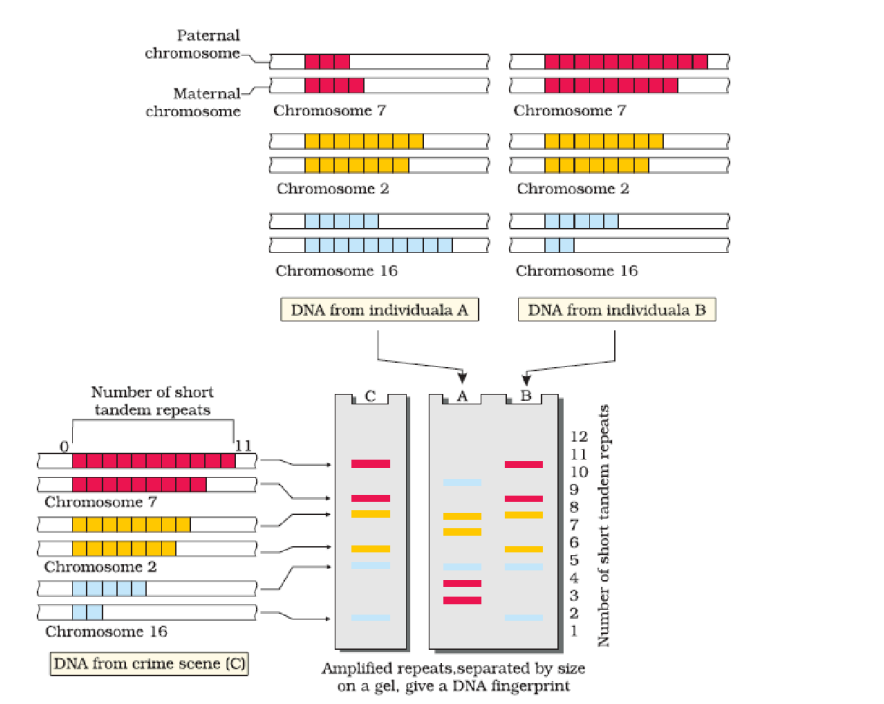
Consequently, DNA from a single cell is enough to perform DNA fingerprinting analysis. In addition to application in forensic science, it has much wider application, such as in determining population and genetic diversities. Currently, many different probes are used to generate DNA fingerprints.
Frequently Asked Questions
It is the study of how genetic information is stored, expressed, and passed from one generation to the next at the molecular level (DNA, RNA, proteins).
- Frederick Griffith (1928): Transformation experiment
- Avery, MacLeod, and McCarty (1944): Proved DNA is genetic material
- Hershey and Chase (1952): Confirmed DNA is genetic material using bacteriophage
- Proposed by Watson and Crick (1953)
- Double helix with sugar-phosphate backbone
- Nitrogen bases: Adenine (A)–Thymine (T), Cytosine (C)–Guanine (G)
- Bases paired by hydrogen bonds
- DNA has deoxyribose sugar, RNA has ribose sugar
- DNA has Thymine, RNA has Uracil instead
- DNA is double-stranded, RNA is mostly single-stranded
The process of copying DNA before cell division. It is semi-conservative, meaning one strand of parent DNA is conserved in each daughter DNA.